February 1: Pattern Formation, Spatial Control of Gene Expression
Please read chapter 4 in the Carlson textbook (pages 58-73 in the fifth edition)
It is a good short summary of the molecular genetic side of embryology.
Concentrate on the principles, rather than the details or the names of the genes.
How Gene Expression (mRNA and protein synthesis) is spatially controlled in embryos.
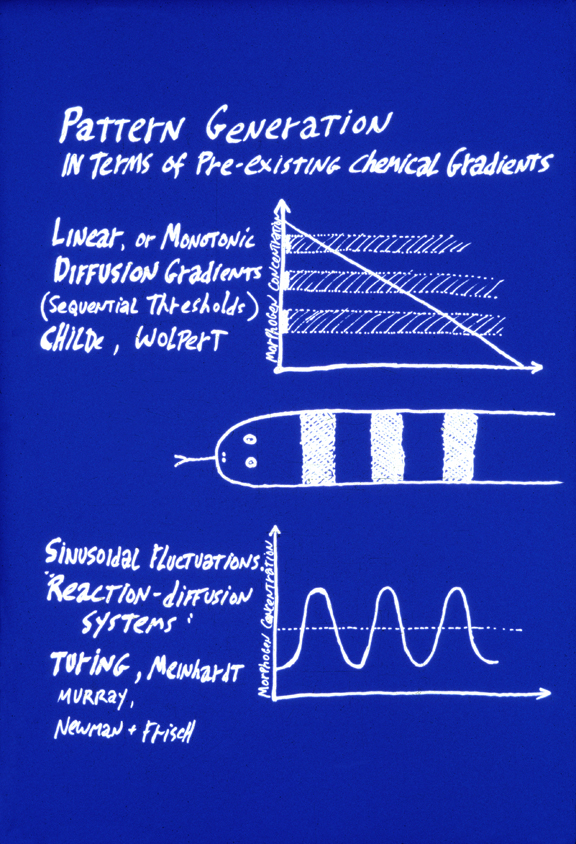
The top of this figure represents the theory of "Positional Information"
This concept was invented by Lewis Wolpert, specifically as a way to explain the Driesch phenomenon. e.g. separated 2 cell stage echinoderms developing into half-size larvae, and all that. "
-
a) Three or more linear diffusion gradients form, each perpendicular to the other two.
b) Each cell measures the three concentrations at its location of these three "morphogen" chemicals.
c) An unspecified molecular mechanism "interprets" the combination of three morphogen concentrations.
(For example, if concentration #1 were 22% maximum, and #2 were 55% maximum and #2 were 77% maximum. then the unspecified interpretation could be "differentiate into a chondrocyte".
The key idea is that diffusion gradients should be twice as steep in half-sized embryos, ten times as steep in tenth-sized embryos, half as steep in double-sized embryos.
** The chemical morphogens of positional information are very different that what Turing invented that word to mean.
*** The spatial variation of morphogen concentrations does not form any geometric pattern (other than linear, or at least monotonic gradients).
**** Most developmental biology textbooks do not distinguish between these uses of the word "morphogen". Sometimes they mean one definition, sometimes they mean the other!
******************************************************
Genes and proteins affecting embryonic pattern formation show extreme (unexpected) conservativeness.
How were such facts discovered? .
Genetic screens: systematic search for mutant animals that are abnormal in interesting ways
For example, you might search for mutant fish that don't swim well, or in which the heart fails to form. .
Looking for needles in haystacks = try using a magnet. .
Genes are named for what goes wrong when the gene is mutated.
Names are often "whimsical" - mutants with no hearts might be called "Heartless", but the name "Tinman" (from The Wizard of Oz) was chosen for one of the best-characterized genes that can be mutated to produce embryos that fail to develop a heart. The function of the non-mutant form of that gene is construction of the heart. .
Many genes that alter embryos code for transcription factors
Families of transcription factors include: Hox, zinc finger, leucine zipper, and others
-----------
These gene sequences are conservative through evolution (for example have homologous sequences in flies and in vertebrates) although they don't necessarily have the same function.
-----------
Five families of Drosophila genes
Segmentation: Gap (Kruppel), Pair-rule (even-skipped) (there is also an odd-skipped mutant, called Fushi Terazu after a kind of Japanese bamboo), Segment-polarity (engrailed)
Homeotic (antennapedia [mutant forms legs on the head where the antennae should be], bithorax [mutant has an extra thoracic segment, and therefore develops an extra pair of wings])
Hox (homeotic) genes - found in "all" animals
From the Wikipedia article on Hox genes:
"A fly can function perfectly well with a chicken Hox protein in place of its own. So, despite having a last common ancestor that lived over 670 million years ago, the chicken and fly version of the same Hox gene can actually take each other's places when swapped." .
"Colinearity of Hox genes
In some organisms, especially vertebrates, the various Hox genes are situated very close to one another on the chromosome in groups or clusters. Interestingly, the order of the genes on the chromosome is the same as the expression of the genes in the developing embryo, with the first gene being expressed in the anterior end of the developing organism. The reason for this colinearity is not yet completely understood ".
"The homeodomain [of a Hox protein] is a 60-amino-acid-long DNA-binding domain (encoded by its corresponding 180-base-pair DNA sequence, the homeobox).... This
consensus polypeptide chain is (typical intron position noted with dashes):
RRRKRTA-YTRYQLLE-LEKEFLF-NRYLTRRRRIELAHSL-NLTERHIKIWFQN-RRMK-WKKEN"
Converting this from the single-letter amino acid code, this protein consists of
Arginine-Arginine-Arginine-Lysine-Arginine-Threonine-Alanine-Tyrosine-Arginine-Tyrosine-Glutamine-Leucine-Leucine-Glutamate-Leucine-Glutamate-Lysine-Glutamate-Phenylalanine-Leucine-Phenylalanine-Asparagine-Arginine-Tyrosine-Leucine-Threonine-Arginine-Arginine-Arginine-Arginine-Isoleucine-Gluamate-Leucine-Alanine-Histidine-Serine-Leucine-Asparagine-Leucine-Threonine-Glutamate-Arginine-Histidine-Lysine-Isoleucine-Tryptophan-Phenylalanine-Glutamine-Asparagine-Arginine-Arginine-Methionine-Lysine-Tryptophan-Lysine-Lysine-Glutamate-Asparagine
Notice how many arginines and lysines there are. Having a lot of these two amino acids makes a protein basic, and therefore able to bind to acidic molecules, i.e. DNA. (This is all you need to remember about this sequence.)
-----------
-----------
-----------
-----------
-----------
**************************************************************************
Another protein that is highly conserved is called Bone Morphogenetic Protein (see this Wikipedia article for more information than was presented in class.
This protein is now being used to induce bone formation, in particular fusion of vertebrae, as an alternative to risky surgery.
Decapentaplegic is the Drosophila gene that is homologous to mammalian BMPs


Maternal effect - establish gradients - bicoid is example




